  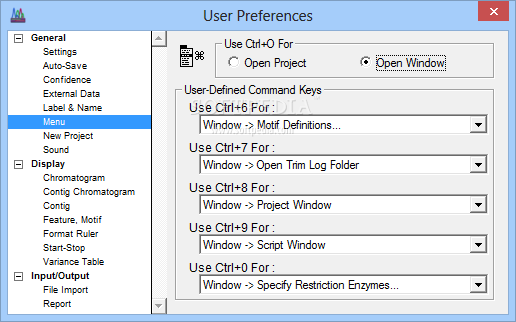
Slow down under certain Windows install due to autosaving Bug in "show info" toggle under Windows and Linux Detection of overlapping Primers in PCR a preference to limit or not the width of Seq & Prot windows (see the available contextual help window to get more details on new functions) It also allows to generate sense and antisense sequences to be obtained after bi-sulfite conversion. Serial Cloner 2.5 now allows to import codon frequency tables from the internet using a Web interface. It is also posible to send directly BLAST request at the NCBI and obtain the result inside a Web interface. You will also find a Restriction Enzyme library management interface, Additional tools, like a web browser for direct import of NCBI and EMBL entries, a virtual cutter to prepare restriction analysis or a silent restriction map generator to find how to introduce restriction sites without modifying the translated peptide are also provided. Features are now visible when aligning locally. 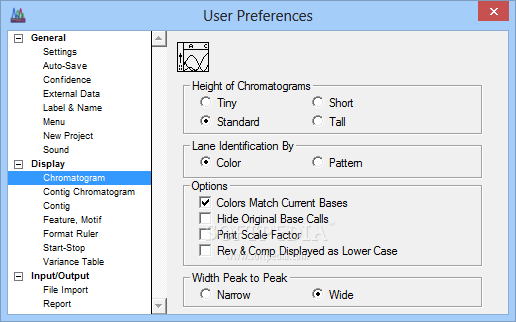
Finally, Serial Cloner provides an interface to align two sequences using a local algorithm or the BLAST2Seq NCBI server. An additional interface allows easy Gateway(tm) cloning for both BP and LR reactions. Just select, blunt if you need, and click the Ligate button. Finally, you can assemble fragments, obtained by PCR, adaptor/shRNA synthesis or simply by graphically selecting fragments between restriction sites. shRNA constructions based on pre-defined scaffolds are also automated. Participation in other Delaware INBRE-supported projects through its Outreach and Administrative Cores such as such as workshop attendance, travel support to conferences, or hosting a Delaware INBRE undergraduate research intern.PCR-based fragment or synthetic adaptors.Bioinformatics resources supported by INBRE include: Vector NTI, Vector Xpression, SeqWeb – GCG, Sequencher, Molecular Operating Environment (MOE), Starlight, Amira Use of bioinformatics resources supported by BRIN/ Delaware INBRE.Use of Core Instrumentation Facilities supported by Delaware INBRE: Bioimaging Center at DBI, Bioinformatics Center at DBI, Microarray Core Center at DBI, Analytical Ultracentrifuge at DBI, Chemi-luminescence Imaging System in Wolf Hall, Protein/Peptide Sequencing facility in Brown Lab.Direct support of research by Delaware INBRE.Following are examples of support through Delaware INBRE that may have made publication of your work possible: Support from Delaware INBRE can come in many ways. If the work that is published was supported in any way by Delaware INBRE, then this support should be acknowledged. This project was supported by the Delaware INBRE program, with a grant from the National Institute of General Medical Sciences – NIGMS (P20 GM103446) from the National Institutes of Health and the State of Delaware.This publication was made possible by the Delaware INBRE program, supported by a grant from the National Institute of General Medical Sciences – NIGMS (P20 GM103446) from the National Institutes of Health and the State of Delaware.These materials must include one of the following two statements: This includes grantees receiving direct support from Delaware INBRE as well as indirect support, such as through the use of Delaware INBRE resources such as the Delaware Biotechnology Institute Core Instrumentation Centers. To that end, all grantee publications – including research publications, articles, abstracts and presentations that are funded by NIH and the National Institute of General Medical Sciences (NIGMS) – require acknowledgement. Publicizing the outcomes of projects that are funded by the National Institutes of Health helps the community better understand the role of biomedical research and its impact on improving human health. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
